|
11/20/2023 0 Comments Flowjo panel analysis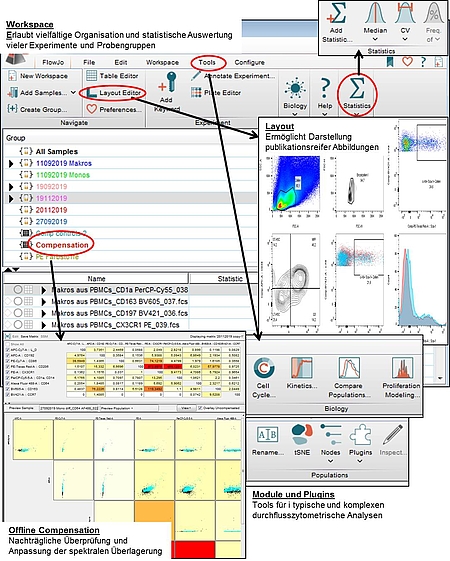
Ortega SB, Torres VO, Latchney SE et al (2020) B cells migrate into remote brain areas and support neurogenesis and functional recovery after focal stroke in mice. Monaco G, Chen H, Poidinger M et al (2016) flowAI: automatic and interactive anomaly discerning tools for flow cytometry data. McInnes L, Healy J, Melville J (2018) UMAP: uniform manifold approximation and projection for dimension reduction. Linderman GC, Rachh M, Hoskins JG et al (2019) Fast interpolation-based t-SNE for improved visualization of single-cell RNA-seq data. Liechti T, Roederer M (2019) OMIP-060: 30-parameter flow cytometry panel to assess T cell effector functions and regulatory T cells. Nat Biotechnol 31:545–552īelkina AC, Ciccolella CO, Anno R et al (2019) Automated optimized parameters for T-distributed stochastic neighbor embedding improve visualization and analysis of large datasets. Key wordsĪmir E-AD, Davis KL, Tadmor MD et al (2013) viSNE enables visualization of high dimensional single-cell data and reveals phenotypic heterogeneity of leukemia. While these methods can identify long-term neuroinflammatory responses after stroke, the methods could be applied to a variety of flow cytometry data sets. The section will describe pre-processing and outline the steps needed to perform unsupervised clustering of your data set in addition to using nonlinear dimensionality reduction for visualizing your analysis. This chapter gives a step-by-step guide on using modern computational approaches to analyze complex flow cytometry data sets in FlowJo™ Software v10. Recent developments have moved the analysis of high-parameter flow cytometry data sets from the traditional analysis method of manual gating to using unbiased analyses to improve scientific rigor. 2017, doi:10.12688/ cytometry has been used for the last two decades to identify which immune cell subsets diapedese from the periphery into the brain parenchyma following injuries, including ischemic and hemorrhagic stroke. “CyTOF workflow: differential discovery in high-throughput high-dimensional cytometry datasets.” F1000Research vol. The following is an example of a published R-based mass cytometry analysis pipeline It’s relatively easy to learn and provides the greatest power and flexibility to analyze mass cytometry data through the packages developed to handle mass cytometry data. R is a programming language with a strong focus on statistics that has become a favorite platform for mass cytometry analysis. Analysis with a programming language R and Python CytobankĬytobank is a web-based software solution that provides t-SNE dimensionality reduction and FlowSOM, SPADE and CITRUS for clustering analysis. *Please let the IMC Platform team know if you are interested in getting a discounted yearly OMIQ license through our platform. It is a cloud-based program capable of performing all stages of CyTOF data analysis. Data analysis cloud-based programs for Helios OMIQ FlowJo offers dimensionality reduction (via t-SNE) in the base package, and the new Plug-in system allows users to utilize a small suite of packages such as FlowSOM, Phenograph and flowClean to analyze high dimensional data. Here you can find an approach to perform data analysis using Pathsetter.ĭata analysis of Maxpar Direct Immune Profiling assay with Pathsetter (PDF, 1.5 MB)įlowJo, a standard tool used for flow cytometry data analysis, is currently expanding into mass cytometry data analysis. It is important to note that this pipeline only works for this kit. Maxpar® Pathsetter™ () is a fully automated reporting and data analysis solution that automatically identifies 37 immune cell types in FCS files from samples processed with the Maxpar Direct™ Immune Profiling System (MDIP kit). Automated single-cell analysis and reporting Maxpar Pathsetter for CyTOF data from Fluidigm Data analysis companies Astrolabe DiagnosticsĪ company offering full CyTOF data analysis for users with no analysis experience or informatics support. We cover this topic and how to clean up your data in the Introduction to Helios course (see Training for more details). The IMC Platform employs the R algorithm Premessa to perform these tasks and highly recommends it to users as well. If the user would like this done as a service, the IMC Platform team can do this for the price of a half hour consultation. Data PreprocessingĪfter acquisition on the Helios, the user is responsible for panel editing, concatenation, normalization (with EQ beads) and debarcoding. At present, the IMC platform is not able to provide the service of data analysis but will assist you in finding a solution. We recommend getting in touch with a data analyst already in the planning phase of your experiments to ensure that the data you generate is of sufficient quality and all necessary controls are measured.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
